If scientists could slip into a microscopic submarine and sail among the body’s cells, as in the 1966 film “Fantastic Voyage,” one of their first stops would surely be the liver. The unique structure of the largest internal organ in our body includes small hexagon-like functional units called lobules, each of which carries out more than 500 different processes simultaneously, making the liver not only a vital organ but one of the most impressive and complex work systems nature has built.
Now, that is possible — virtually, at least. In a new study recently published in the scientific journal Nature, scientists from the Weizmann Institute of Science, together with colleagues at Sheba Medical Center and the Mayo Clinic in the United States, present for the first time a genetic atlas of the healthy human liver at a resolution of 2 microns. The findings reveal that the division of labor in the human liver differs from that of other mammals and is more extensive than previously known, and show how it makes certain areas of our liver especially vulnerable to fatty liver disease.
6 View gallery

The human liver carries out more than 500 different processes simultaneously
(Illustration: Shutterstock)
“The liver is the largest internal organ in the human body, and it is responsible for many functions,” explains Dr. Oran Yakubovsky, a senior general surgery resident at Sheba Medical Center and a doctoral student at the Weizmann Institute, who was part of the research team. “The liver is responsible for the body’s metabolic activity, heat production, protein production, the coagulation system and filtering toxins — it knows how to take drugs and toxins from the blood, change their form and remove them from the body. It is also responsible for the body’s sugar regulation together with the pancreas.”
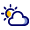
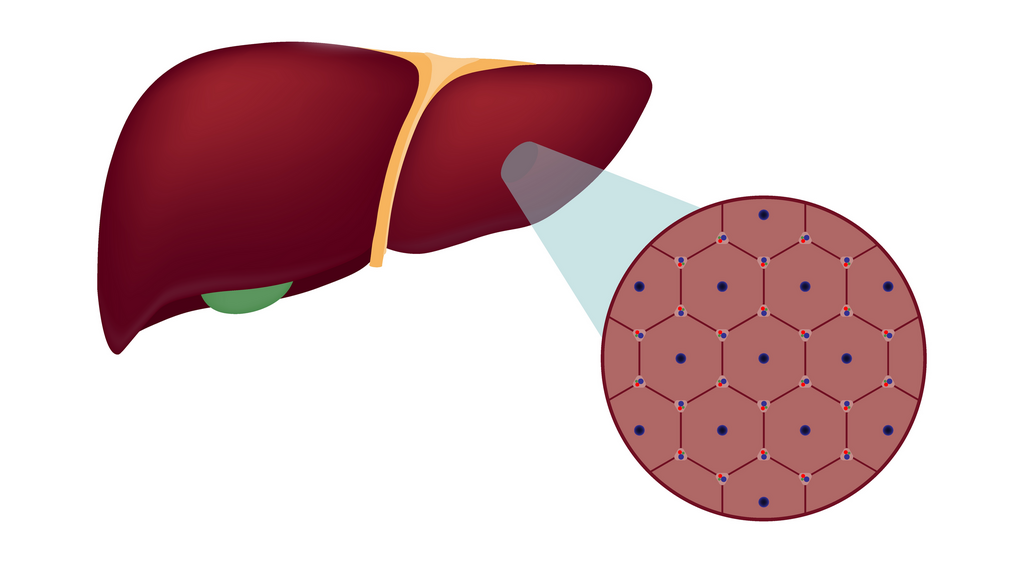
That complexity does not occur on a large scale, but rather at an especially tiny resolution. “The liver is built from very small functional units called lobules,” he adds. “Blood flows from the digestive system into the liver and enters the lobule, which looks like a hexagon, through small blood vessels located around its perimeter. From there it flows inward to the center, and from there it flows to the rest of the body. Inside this lobule, all the actions of liver cells on the blood are actually carried out.”
Since the 1970s and 1980s, it has been known that liver cells divide their many tasks according to their location within each subunit, but the technology at the time allowed only a blurred picture of that division. “They would simply take a liver sample, cut it and examine it under a microscope,” Yakubovsky says.
At the time, researchers also found that the lobule had several distinct regions. “In the outer part, the blood is very rich in nutrients and oxygen, and the farther inward it moves, the more those nutrients are used by the liver cells, so the blood that reaches the center of the lobule is used blood, poorer blood,” he continues. “As a result, a difference can be seen between the outer and inner parts. That was known. Because of that, until now it was customary to divide the lobule into three different zones — from the outside inward.”
Innovative 'atlas'
In recent years, technology has developed that makes it possible to identify which genes are active in each individual cell while locating the cells precisely in space on the tissue map. Still, despite these new capabilities, precise mapping of the division of labor in the human liver was delayed, in large part because of the difficulty of obtaining liver samples from healthy people. In Professor Shalev Itzkovitz’s lab at the Weizmann Institute, researchers realized that the key to the solution was altruistic liver donations: The liver has a remarkable ability to regenerate, enabling people to donate a large portion of it to someone suffering from disease. With the help of Professor Ido Nachmany and Professor Niv Pencovich from Sheba Medical Center’s general surgery department and Dr. Timucin Taner of the Mayo Clinic transplant center in Minnesota, the scientists obtained eight samples from healthy donors and assembled an “atlas” of genetic expression in the human liver.
6 View gallery
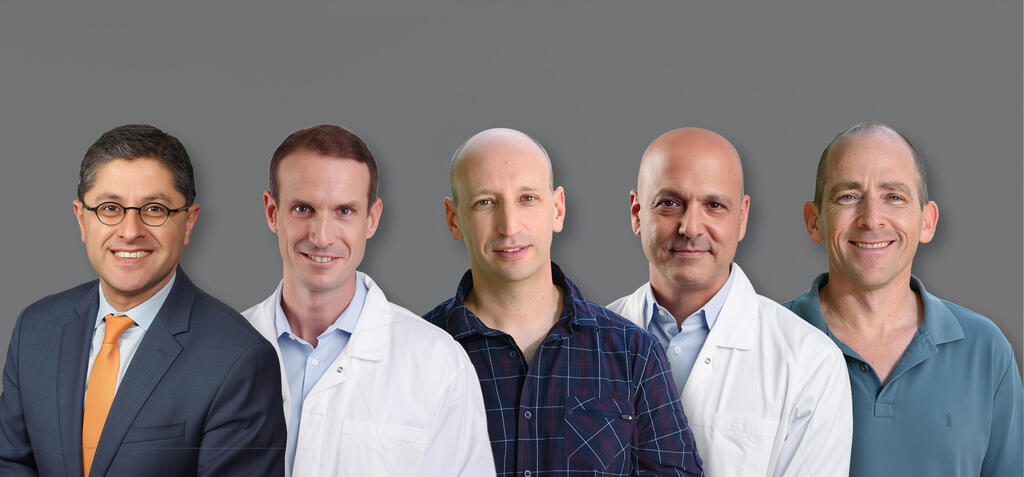


From right, Prof. Shalev Itzkovitz, Prof. Ido Nachmany, Dr. Oran Yakubovsky, Prof. Niv Pencovich and Dr. Timucin Taner
“Until just a few years ago, to characterize what a cell does, you had to look at it under a microscope, stain it, see what it responds to and how it behaves. But since the Human Genome Project, there are advanced methods that allow us to read genetic material in the cell called RNA,” Yakubovsky continues. “That allows us to characterize it at a deeper level — what proteins are there, what it knows how to do. That sequencing really brought about a revolution. If we once thought that every liver cell performs the same functions, today we can say that there are many types of cells doing different jobs depending on their location within the lobule.”
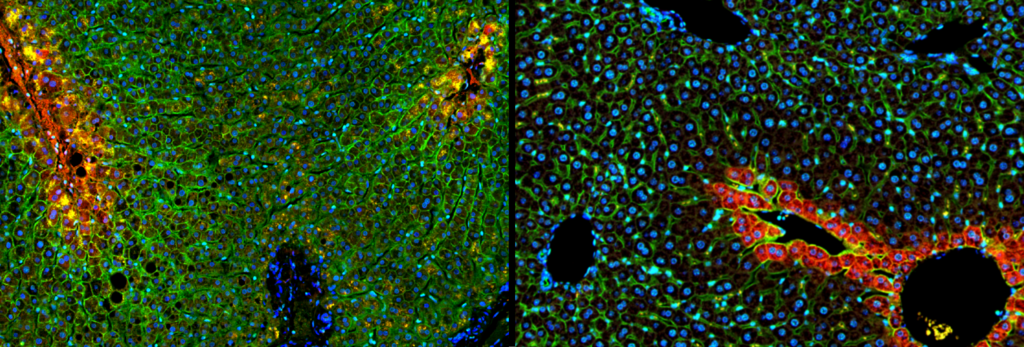
In fact, the discovery allowed researchers for the first time to see the liver at a resolution that had not previously been available — not just as one organ, but as a complex system of different regions, each functioning in a unique way. “Thousands of genes were found to be active at different levels in liver cells in different locations, indicating a far more precise and complex internal organization than we had thought,” Itzkovitz says.
“Instead of the rough division into three activity zones, which has been accepted for decades, the atlas revealed eight regions with distinct roles. The precise mapping of the liver now allows any laboratory in the world to dive deep into the liver and study why different regions are vulnerable to different diseases. Metabolic diseases, for example, tend to begin at the center of the lobule, while viral and autoimmune inflammations appear mainly at its edges. Liver cancer and metastases from other cancers also have their preferred locations. The key to understanding why this is the case lies in the precise genetic information we collected.”
This breakthrough emerged not only from advanced technology, but also from a major weakness in the research: Until now, scientists simply did not have a reliable map of a healthy human liver for comparison. “Until now, we did not have an ‘atlas’ of a healthy human liver that could serve as a healthy reference to work from, like a computerized map book,” Yakubovsky confirms. “It is important to have a reference of a healthy liver from which you can go on to study all the diseases.”
This breakthrough is unique because until now, all the livers described in the scientific literature did not come from healthy donors. “All the livers in the literature are from donors who suffered brain death and donated their organs, but brain death can also affect the liver. Other livers used came from people who underwent surgery to remove a liver tumor, and in those operations a sample was taken from a healthy area near the tumor. We challenged the existing perception and argued that these two methods are not precise enough. We are the first to characterize the human liver using healthy samples, and the only way to obtain that is from liver donors. Now, for example, anyone studying viruses anywhere in the world can enter our atlas and try to search where their virus will have the greatest effect.”
The human liver works differently
To compare humans with other species, Itzkovitz’s lab also mapped the healthy liver in mice as well as in larger mammals — pigs and bulls — whose metabolic rate and liver lobule size resemble those of humans. As noted, in all mammals, blood flows through the lobule from the margins to the center, supplying oxygen and nutrients to cells along the way. As a result of this route, conditions at the margins are abundant, while the center is relatively deprived. In all mammals examined, except humans, the scarcity conditions at the center of the lobule caused cells there to be relatively less active. In humans, however, many processes were found to take place in the core of the lobule, including fat production from excess energy, producing sugar from non-carbohydrate sources during fasting, filtering toxins and producing bile, which aids digestion.
Another significant difference found in the study between the human liver and that of other mammals involves sugar storage. The liver functions as the body’s “fuel tank” by efficiently absorbing the sugars we digest during meals and releasing them in a controlled manner between meals. The study found that in humans glucose absorption is unique to the centers of the lobules rather than their margins, as in mice.
“This division of labor is both a blessing and a curse,” Itzkovitz explains. “It allows our liver to store carbohydrates efficiently: Cells at the center of the lobule absorb and store sugar (glucose) directly from the blood, while cells at the margins convert lactate into glucose and thus also contribute to the energy reserves we use while fasting. However, this efficient division of labor was not designed for the modern diet, which is rich in fats and carbohydrates, and may explain why we tend to accumulate excess fat at the center of the liver lobule and suffer from its scarring.”
To cope with wear and prevent disease, a unique turnover mechanism developed at the center of the human liver lobule. “We found that in humans, unlike other mammals, one type of immune cell prefers the core of the lobule rather than standing guard at its margins — the gateway through which blood enters the tissue,” Yakubovsky says. “Kupffer cells are phagocytic cells that can protect against infections but also engulf, break down and recycle worn-out cell remnants. We assume they ‘moved to the center’ in humans to cope with the increased wear.”
This comparison is of central importance, since much medical research is based on animal models, and differences between them and humans can affect the interpretation of findings. “This is important because around the world, researchers usually do not work on human tissue, which is very difficult to obtain. They mainly work on mice, pigs or other mammals,” Yakubovsky adds. “For example, if a pharmaceutical company is developing a drug and testing it on a mouse model, it can now check our atlas and see where the genes they are working on are expressed, both in mice and in humans.”
How fatty liver develops from within
In the final part of the study, the scientists showed how the new atlas helps trace the development of diseases. They focused on metabolic fatty liver disease, a common condition linked to excess weight and diabetes but found in varying degrees in about 25% of the global population, in which fat accumulates in the liver and can lead to inflammation and scarring. This was made possible thanks to the special collaboration with the Mayo Clinic transplant center in Minnesota. “We took healthy liver samples from human donors — people who underwent a very significant evaluation and were proven to be healthy. But because of the prevalence of fatty liver disease, some of the donors showed early stages of fatty liver,” according to Yakubovsky.
In other words, the uniqueness of these samples allowed the researchers to look not only at advanced disease, but also at its starting point. “Until now, many places that studied fatty liver disease studied advanced fatty liver,” Yakubovsky continues. “We worked with samples from people who had fatty liver at an early stage that was still considered healthy. That is how we were able to capture the earliest stages of the disease and characterize the genes that are expressed.”
The researchers compared healthy liver cells with cells that had begun accumulating fat and discovered several surprising findings. “We found a defense mechanism,” Yakubovsky says. “Cells that had begun to become fatty essentially turned off genes that cause fat uptake and activated other genes that cause its breakdown. In other words, the liver really tries to protect itself in the early stages from that fat absorption inside the liver cells.” However, they also found that the human liver has a failure that limits its ability to effectively resist fat accumulation: Fat buildup was found to lead to a decline in the production of some components in mitochondria — organelles that break down fats.
After years in which medicine viewed the liver as a single organ, the new map reveals a far more complex reality, one that may also change the way we try to repair it. “Based on the precise mapping of the liver, it will be possible in the future to develop treatments focused on the genes that cause a certain region to be especially vulnerable to a specific disease,” Itzkovitz says. “Moreover, the model of building a genetic atlas at single-cell resolution from samples of healthy donors can be applied to additional organs that until now have not been precisely mapped in humans, and it may fundamentally change the way we understand the structure and function of the human body.”
After all the layers of complexity, the comparisons between species and the attempt to understand exactly where disease begins, the researchers also want to leave behind something more basic: infrastructure. One that can now serve other researchers anywhere in the world as a reliable reference point for studying the human liver. “We hope this foundation will become the reference for all liver work going forward, meaning that any lab or researcher that wants to develop drugs or examine certain diseases will use our atlas to check their results,” Yakubovsky says. “From drug development to characterizing specific liver diseases, our atlas will serve as a starting point for research.”
The study also included Dr. Keren Bahar Halpern, Sapir Shir, Roy Novoselsky, Dr. Adi Egozi, Dr. Tal Barkai, Dr. Yotam Harnik, Dr. Amichay Afriat and Dr. Yael Korem Kohanim of the Weizmann Institute’s Department of Molecular Cell Biology; Dr. Chen Meir and Dr. Ron Perry of Sheba Medical Center at Tel Hashomer; Dr. Ruben Hoflin of the University of Freiburg in Germany; Ofra Golani, Dr. Ina Goliand, Dr. Yosef Addi, Dr. Meirav Kedmi and Dr. Hadas Keren Shaul of the Weizmann Institute’s Life Sciences Core Facilities Department; and Dr. Liat Plus-Aligor, Yelena Pritchislov and Dana Hirsch of the institute’s Veterinary Resources Department